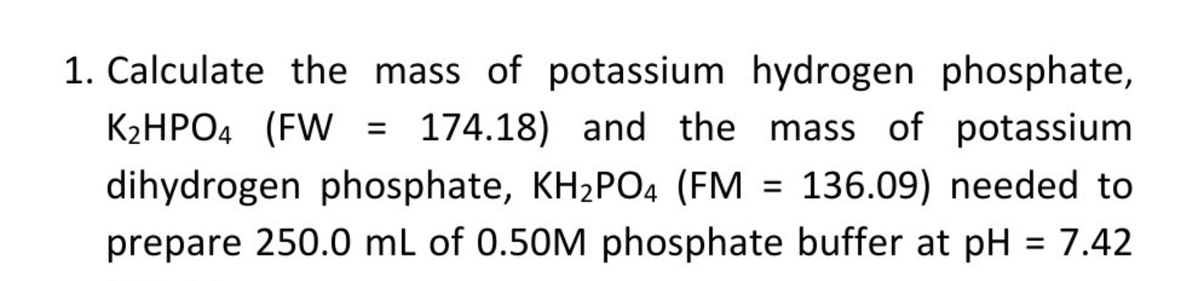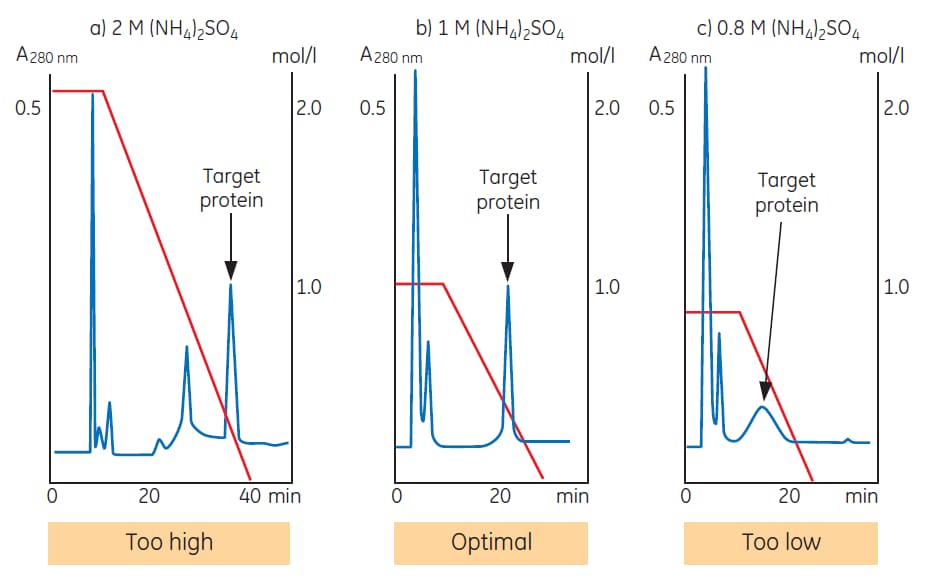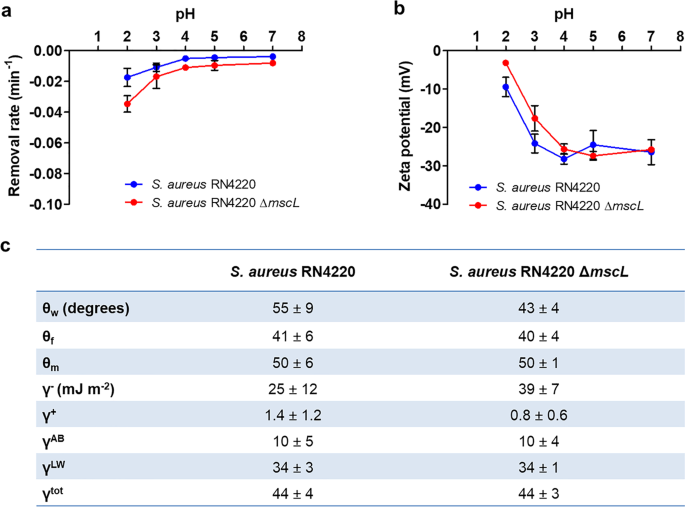
Role of adhesion forces in mechanosensitive channel gating in Staphylococcus aureus adhering to surfaces | npj Biofilms and Microbiomes

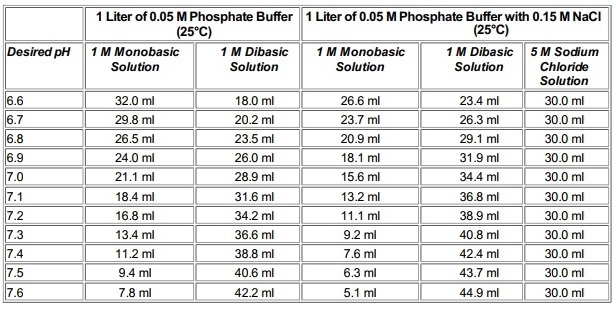
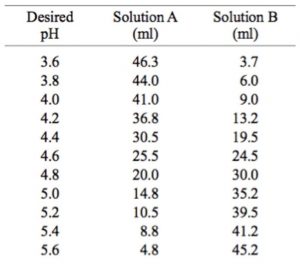
Table 2 from Dynamic approach to predict pH profiles of biologically relevant buffers | Semantic Scholar
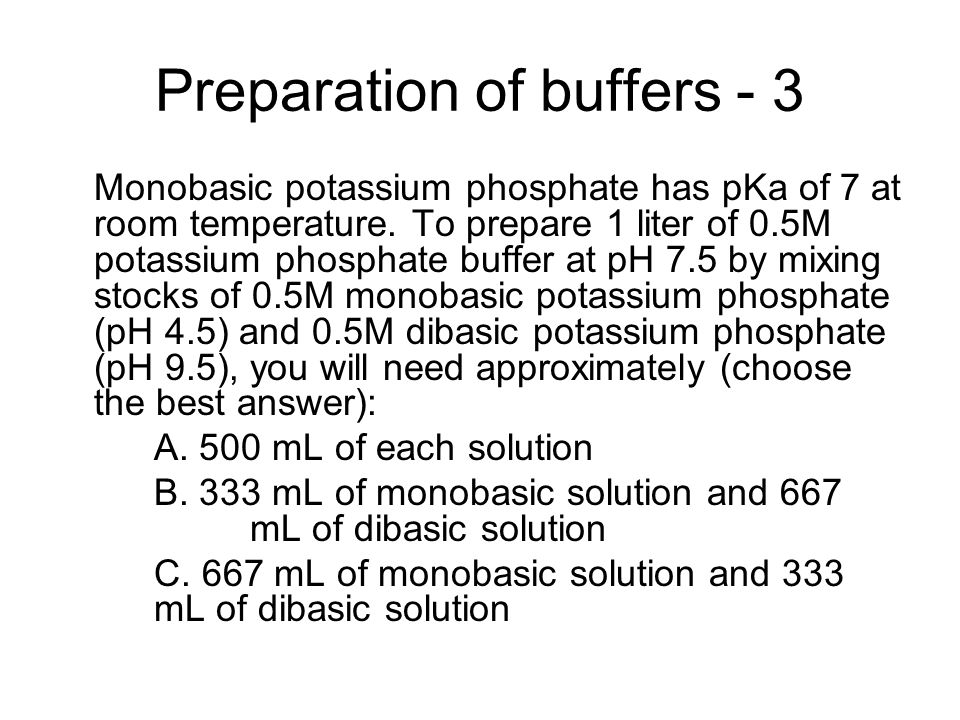

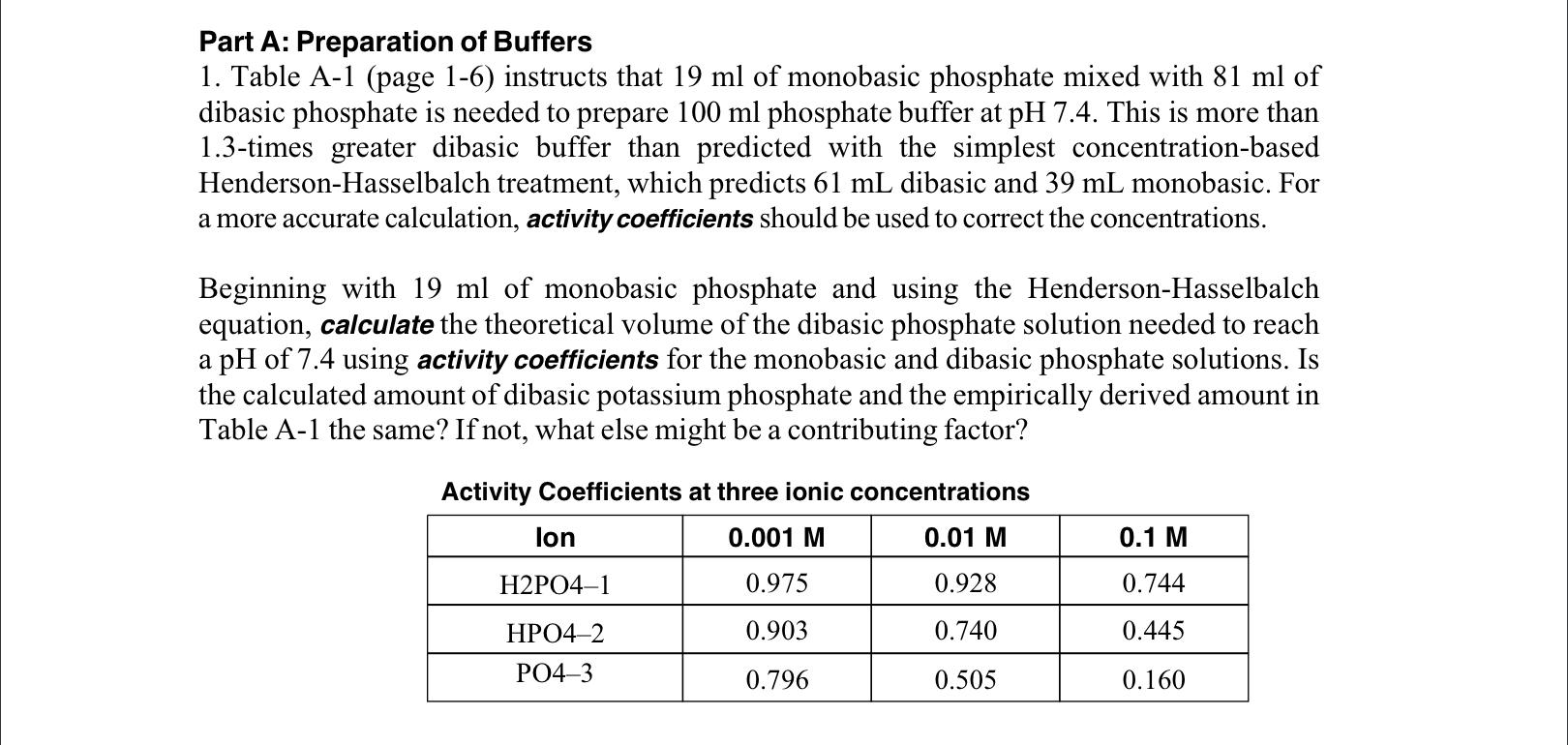
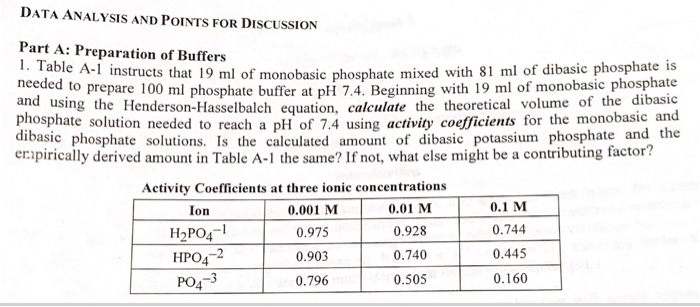
Preparation of Buffers - 1 Calculate the volume of sulfuric acid (H 2 SO 4 ) necessary to prepare 600 milliliter 0.5M H 2 SO 4 from concentrated H 2 SO. - ppt download
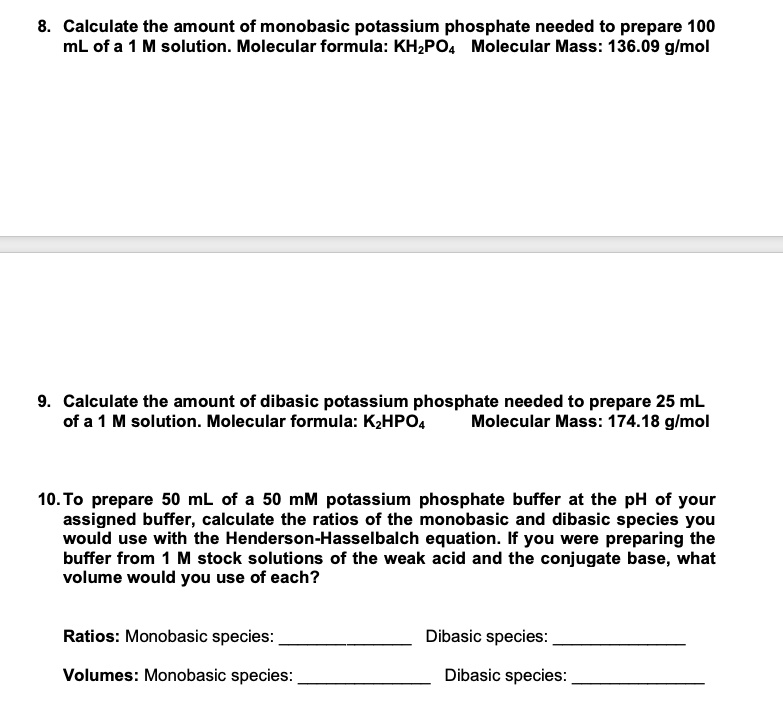

SOLVED: Calculate the amount of monobasic potassium phosphate needed to prepare 100 mL ofa M solution Molecular formula: KHZPOa Molecular Mass: 136.09 glmol Calculate the amount of dibasic potassium phosphate needed to
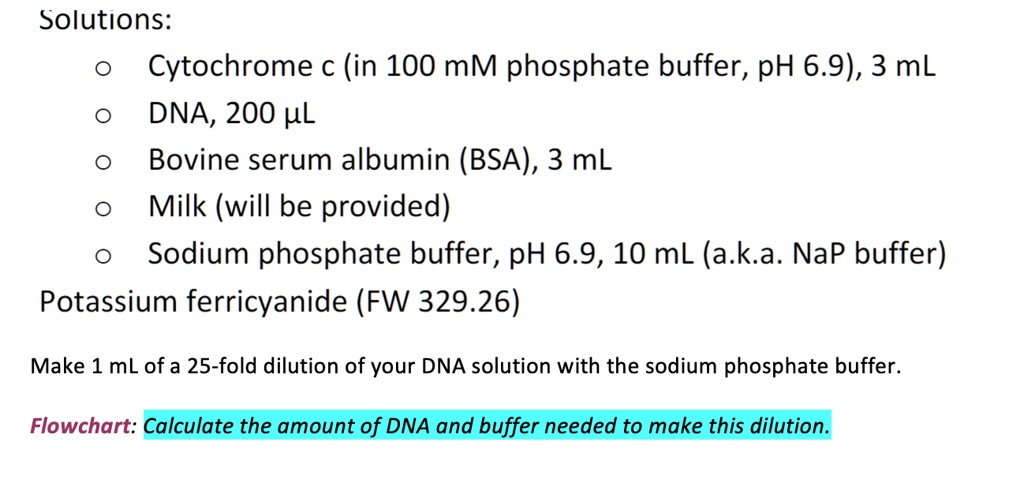

SOLVED: Solutions: Cytochrome c (in 100 mM phosphate buffer, pH 6.9),3 mL DNA, 200 HL Bovine serum albumin (BSA), 3 mL Milk (will be provided) Sodium phosphate buffer, pH 6.9, 10 mL (

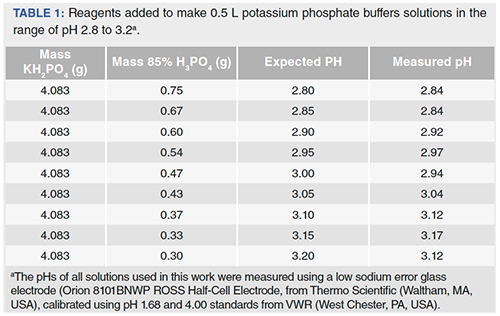
Table 1 from Dynamic approach to predict pH profiles of biologically relevant buffers | Semantic Scholar